Beet western yellows virus (BWYV): Aspect of Outbreak and Survey, and First Complete Genome Sequence of a Korea Isolate of BWYV
Article information
Abstract
In 2010, foliar symptoms were observed in the paprika leaves in Jinju city, Korea. Beet western yellows virus (BWYV) was identified in paprika by using the large-scale oligonucleotide chip assay. To investigate the occurrence of BWYV, a survey was performed on various crops, including paprika, from 2011 to 2014. Further, the presence of BWYV was consistently verified through literature survey from 2015 to 2017. BWYV infection has been identified in Solanaceae crops (bell pepper, hot pepper, and paprika), various weeds, and green peach aphids and it occurs on a nationwide scale. Cultivation using organic methods involved natural enemies and showed a high BWYV infection rate, which was more than that for conventional cultivation methods in greenhouse. The complete genome sequence of BWYV isolated from paprika was determined for the first time. The genome of the BWYV-Korea isolate consists of 5750 nucleotides and has six open reading frames. Sequence identity results showed maximum similarity between the BWYV-Korea isolate and the BWYV LS isolate (identity > 90%). This study is the first report of BWYV infecting paprika in Korea. The survey revealed that BWYV is naturalized in the domestic ecology of Korea.
Introduction
According to the International Committee on Taxonomy of Viruses (ICTV; http://www.ictvonline.org), 17 species are currently approved as members of the genus Polerovirus. Of them, Beet western yellows virus (BWYV), belonging to the family Luteoviridae, was first reported in Beta vulgaris, Lactuca sativa, Spinacia oleracea, and Raphanus sativus by Duffus (1960, 1961) in California, USA. BWYV has a broad natural host range, infecting more than 150 species in 23 dicotyledonous families (Brunt et al., 1996). This virus was reported in Asia (China, Israel) (Marco, 1984; Zhou et al., 2011), Europe (Czechoslovakia, England, Scotland, Denmark, Germany) (Pálak, 1979; Russell and Duffus, 1970), North America (USA) (Beuve et al., 2008), and Oceania (Western Australia) (Hertel et al., 2004). It is obligately transmitted by one or a few aphid species in a persistent and circulative mode and causes serious economic losses by reducing crop production (Brault et al., 2005; Brown, 2000; Tamaki and Fox, 1982). This virus can infect phloem cells and consequently cause viral symptoms, such as stunting and reddening of plant leaves (Reinbold et al., 2001).
In general, the genus Polerovirus has a positive-sense single-stranded linear RNA genome of approximately 5.6-6.0 kilobases (kb). BWYV virions are icosahedral particles of 25 nm diameter (Knierim et al., 2014). BWYV genome is organized in six open reading frames (ORFs, ORF0-5) with translational products named as P0-5 (Mayo and Ziegler-Graff, 1996). The ORFs encode the following: ORF0 (P0 protein: host range, symptom expression, and suppressor functions), ORF1 and ORF2 (P1 and P1-P2 fusion protein: viral RNA-dependent RNA polymerase domain), ORF3 (P3 protein: coat protein), ORF4 (P4 protein: movement protein), and ORF5 (P5 protein: aphid transmission-related protein). The ORF2 synthesizes the P1-P2 fusion protein by ribosomal frameshift, and the P3-P5 fusion protein is produced by read-through at stop codon of the P3 protein (Domier, 2012). The 5’ untranslated region (UTR) is covalently linked to the genome-linked viral protein (VPg), and the 3’ UTR has no poly (A) structure (Dreher and Miller, 2006).
BWYV is designated as a plant quarantine virus in Korea, and it was reported for the first time in paprika (Capsicum annuum var. angulosum) by Park et al. (2011). In February 2012, emergency control was carried out by the Rural Development Administration (RDA), after which additional investigations on this virus have not been conducted. Until now, there is no report on the occurrence of BWYV in Korea. Here, we describe for the first time the occurrence of BWYV in paprika in the greenhouses of Korea and its identification by complete genome sequencing of the isolated virus.
Materials and Methods
Plant material and virus detection
Clinical diagnosis of a disease in paprika plants was requested in October 2010 by the Animal and Plant Quarantine Agency (previous name: NPQS; current name: QIA). The foliar symptoms of vein yellowing and vein clearing were observed in the paprika samples collected by Dr. Shin Young-gil in Jinju, Korea.
In order to identify the causal agent, total RNA was extracted from the symptomatic paprika leaf samples using Easy-spin Total RNA Extraction Kit (iNtRON Biotechnology, Daejeon, Korea). Each sample was tested using SuPrimeScript RT-PCR premix (GenetBio, Daejeon, Korea) with six species-specific primers for the major Solanaceae-infecting viruses (Broad bean wilt virus 2, BBWV-2; Cucumber mosaic virus, CMV; Pepper mottle virus, PepMoV; Pepper mild mottle virus, PMMoV; Potato virus Y, PVY and Tomato spotted wilt virus, TSWV) (Choi et al., 2005; Kim et al., 2002, 2004; Mun et al., 2008) (Table 1). In addition, the samples were analyzed by transmission electron microscopy (TEM). After reverse transcription-PCR (RT-PCR) testing, we applied the sample to a large-scale oligonucleotide chip (LSON chip) that contained about four thousand probes to detect five hundred plant viruses and viroids of plant samples (Nam et al., 2014). The chip indicated that the sample was positive for two virus species, and we designed specific primer sets based on the probe sequence. Using these primer sets, we identified the virus infecting the paprika plants; the PCR products were cloned and sequenced. The raw data were edited to eliminate erroneous nucleotide sequences using DNAMAN software Version 7.0 (Lynnon BioSoft, Vaudreuil, Quebec, Canada), and the sequences were identified by NCBI BLASTN search. Further, this sample was used for complete genome sequencing.
Field screening and sample collection
To confirm the occurrence of BWYV, a survey was conducted at three paprika export complexes in Daegok-myeon in Jinju, Gyeongsangnam-do province, Korea. Various crops, such as strawberries, pumpkins, mini paprikas, and hot peppers are cultivated in Daegok-myeon. A greenhouse-based cropgrowing area is also present in Jinju. On 7-8 December 2010, we visited the site to observe the symptoms and incidence of BWYV infection in four farm greenhouses. Each farm cultivated plants using conventional or organic methods, and there were many aphids on symptomatic leaves. In the first investigation, thirteen symptomatic paprika leaf samples were collected (Fig. 1A to C) and two symptomless weed samples were collected from a nearby greenhouse (Fig. 1D and E). On 10-14 January 2011, a second investigation was performed to determine whether BWYV has spread to other host plants. Seventeen paprika leaf samples were collected from the greenhouses of six paprika farms; additional 48 samples were collected from weeds, sweet pepper, curled mallow, and squash plants in and around the greenhouses. In addition, nine green peach aphids (Myzus persicae) were collected from the paprika leaves (Fig. 1F). On 18 February 2014, a third investigation was conducted to confirm the existence of this virus. Eight paprika leaf samples, showing the same symptoms, were collected from the same greenhouse where the first BWYV infection was discovered. All the samples were subjected to RT-PCR, and the nucleotide sequences of positive amplification products were determined by cloning, followed by sequencing.
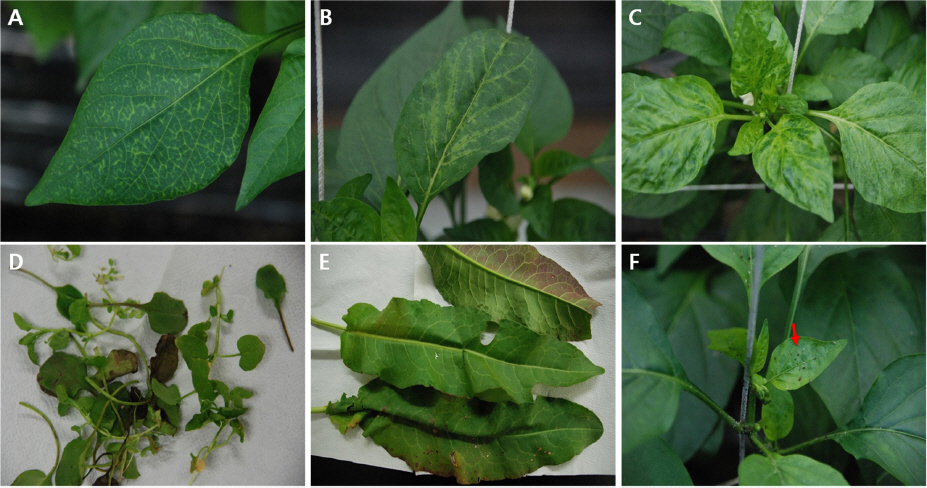

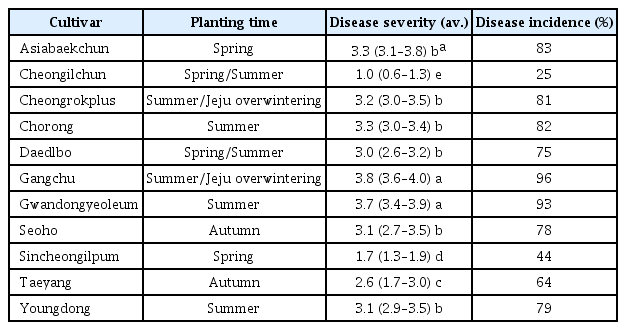
Photographs of plants and aphids infected with Beet western yellows virus (BWYV). Symptoms of BWYV infection in paprika from a farmer’s greenhouse; (A) vein yellowing symptoms, (B) vein chlorosis symptoms, (C) mosaic symptoms. Two weeds samples collected in a nearby greenhouse that were infected with BWYV and showed no symptoms; (D) Trigonotis peduncularis, (E) Rumex japonicus. Numerous aphids (Myzus persicae) observed in the greenhouse where BWYV occurred (F).
Complete genome sequencing and sequence analysis
To determine the complete nucleotide sequence of the paprika-infecting BWYV, four primer pairs were designed based on sequences from published BWYV isolates (Table 2). The cDNA was synthesized with Random N(25) primers using Reverse Transcriptase AMV (Roche, Mannheim, Germany) according to the manufacturer’s protocol. The PCR amplification reaction was conducted in a total volume of 20 μl, which contained 1 μl of cDNA template, 1 μl of each forward primer, 1 μl of each reverse primer, 1 μl of Elongase Mix, 4 μl of 5×Buffer A [300 mM Tris-SO4, (pH 9.1 at 25°C), 90 mM (NH4)2SO4, and 5 mM MgSO4], and 12 μl of RNase-free water (Invitrogen, Carlsbad, CA, USA). The PCR cycle conditions were programmed as follows: pre-amplification denaturation at 94°C for 30 s and 28 cycles of 94°C for 30 s, 55°C for 30 s, and 68°C for 1 min 40 s. The amplified PCR fragments were purified with Expin PCR SV (GeneAll Biotechnology, Seoul, Korea); the purified PCR fragments were confirmed by agarose gel electrophoresis and cloned into a pGEM-T Easy vector (Promega, Fitchburg, WI, USA) using T4 ligase (TaKaRa, Tokyo, Japan). The clone was transformed into competent DH5α cells (RBC Bioscience, New Taipei City, Taiwan) following the manufacturer’s instructions. Eight white clones were identified by PCR premix (GenetBio, Daejeon, Korea) with vector-specific universal primers T7/SP6. Four selected clones were cultured on SOC medium with ampicillin, and plasmid DNA was extracted using Enzynomics plasmid kit (Enzynomics, Daejeon, Korea). The eluted plasmid DNA was used for HindIII restriction enzyme digestion and sequencing.

Oligonucleotide sequences of primers used for complete genome sequencing of Beet western yellows virus (BWYV)-Korea isolate
To determine the 5’ UTR region of the BWYV genome, specific reverse primers (BWYV-R0: GGC CAA ACG GTG ACG TTG GCG & BWYV-R1: CCT GCA ACG TCT TGG GTG) were designed and amplified using 5’ RACE System for Rapid Amplification of cDNA Ends, Version 2.0 (Invitrogen, Carlsbad, CA, USA). To determine the 3’ UTR, specific forward primers (BWYV-F0: TAA CGT TTT TCT TGC TCG CCT & BWYV-F1: AGA CGT TGC AGG ATT ATT CCT) were designed and A-tailing performed using a Poly (A) Polymerase Tailing Kit (Epicentre BIO Technologies, Madison, WI, USA). After A-tailing, cDNA synthesis, PCR, and cloning were carried out as described above. All plasmid and RACE fragment sequences were determined in both directions by Solgent Co. (Daejeon, Korea). The resulting nucleotide sequence was assembled to form a consensus sequence and was edited using DNAMAN software. A phylogenetic tree was constructed using the neighbor-joining method, and nucleotide identities were calculated based on the observed divergences.
Results and Discussion
Identification of BWYV
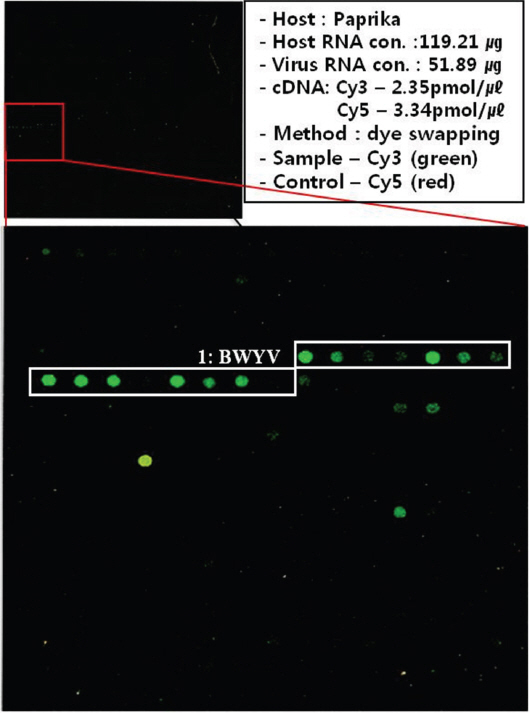
The etiology of the foliar symptoms in paprika plants was confirmed using two detection methods (RT-PCR and TEM). The RT-PCR results were negative for the six known viruses (BBWV-2, CMV, PepMoV, PMMoV, PVY, and TSWV), and virion particles were also not observed in the crude sap. To identify the infectious agent in the visually symptomatic leaf tissue, total RNA was isolated and applied to the LSON chip, and the specific BWYV and CABYV probes showed positive reactions (Fig. 2). To confirm this result, the two viruses were targeted using RT-PCR with specific primers. The RT-PCR results were positive only for BWYV. BLAST search results showed that the nucleotide sequence of this virus was most similar to a BWYV USA isolate (91% identity); thus, we named this virus the BWYV Korea (Kor) isolate.
Results of the field survey
To further investigate the incidence of BWYV, a survey was carried out in 2010, 2011, and 2012 in paprika export complexes in Jinju. In the first investigation, paprika leaves showed virus-like symptoms, such as vein yellowing, vein banding, and mosaic, while two weeds samples did not show any symptoms. All the 13 collected samples were tested by RT-PCR using BWYV 95F/784R species-specific primers. The RT-PCR results yielded an amplicon of 690 bp that was detected in eight paprika and two weed samples infected with BWYV. In addition, three samples were positive for PepMoV. In the second investigation, leaves from various plants were collected, including weeds, bell peppers, and paprika. Seven paprika samples, two sweet pepper plants, and three green peach aphids were positive for BWYV. The nucleotide sequences of the positive samples of one paprika plant, one aphid, and two weeds were aligned, which showed 98.77% nucleotide identity (Fig. 3). These results suggest that each infected plant was associated with green peach aphids in these regions. However, other weeds such as Japanese dock, keijunmekko, curled dock, curled mallow, and squash were negative for BWYV. In the third investigation, eight samples were collected from the same greenhouse where the first BWYV was discovered. The collected samples were tested to confirm the presence of BWYV using specific primer sets. However, all samples were negative for BWYV. Thus, we tested the samples with eight primer sets specific for the other major Solanaceae-infecting viruses. The test results showed that eight samples were positive for PepMoV. To date, BWYV has been reported in various crops and weeds. The BWYV-infected plants showed typical symptoms such as yellowing, oranging, dwarfing, violetting, mild yellowing, mosaicism, and crinkling on the leaves of various plants (Duffus, 1960, 1961; Pálak, 1979; Smith and Hinckes, 1985; Wang et al., 2013; Yuan et al., 2015). However, the BWYV infecting the paprika crops has not yet been characterized. In the second investigation, vein yellowing was commonly detected in plants infected with BWYV. However, the last investigation suggested that vein yellowing symptoms in paprika leaves are not representative of BWYV infection. To verify this result, one positive sample of PepMoV is used as inoculum to inoculate three paprika cultivars (Mazzona, Veyron, and Coletti), which are mainly cultivated in Daegokmyeon in Jinju. At 3-4 weeks post-inoculation, ‘Veyron (red)’ and ‘(Yellow) coletti’ cultivars showed vein clearing symptoms on upper leaves, similar to BWYV infected paprika leaves in the greenhouse. Mazzana (orange) cultivar showed vein banding and mosaic symptoms on upper leaves. Infectious agents in these test plants were confirmed with RT-PCR using PepMoV-specific primer pairs. Our biological test verified that the vein clearing symptom in paprika plants is not because of BWYV infection.
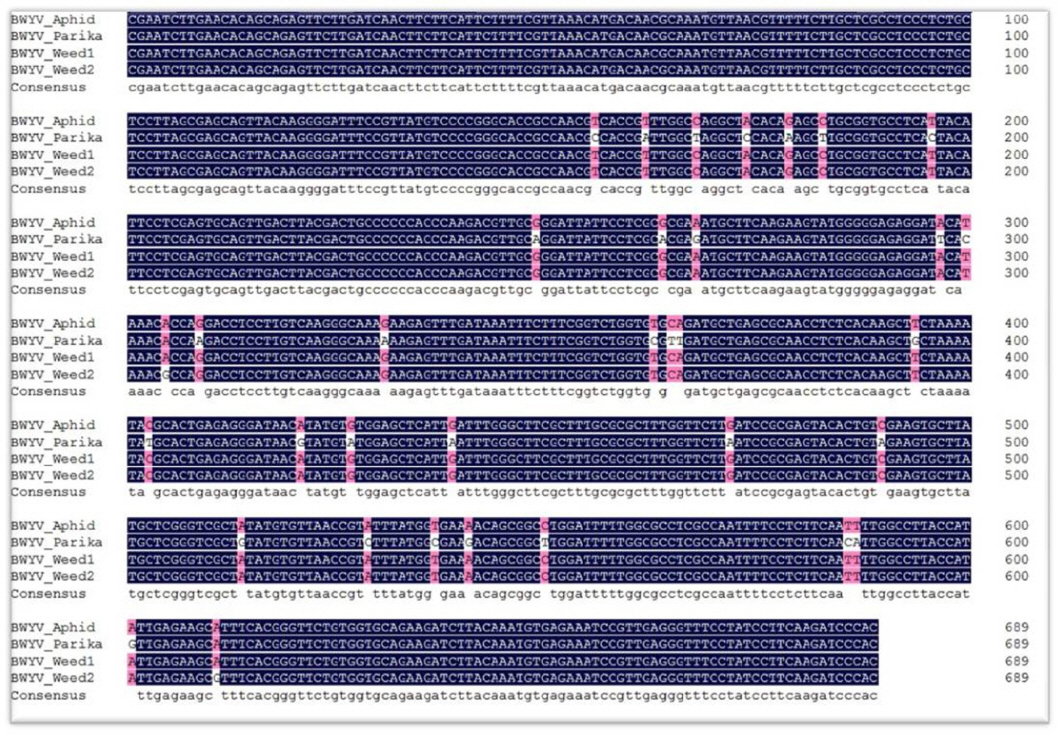
Alignment of the RdRP partial sequences of four samples, including paprika. The nucleotide sequence corresponding to the RdRP region is highly conserved and is marked in blue (Identity: 98.77%).
BWYV was detected in weed species and a green peach aphid, implying that this virus is transmitted to other plants by aphids. Aphids are mainly observed in greenhouses that use organic farming methods; moreover, these greenhouses have been confirmed to have high BWYV infection rates. In order to reduce the spread of BWYV, weeds and aphids must be removed from the greenhouse. In 2014, a nationwide survey (5 Provinces, 13 regions) was conducted by the National Institute of Agricultural Sciences and each region’s Agricultural Research & Extension Services center to investigate the outbreak. According to the reports, BWYV was detected in nine out of the thirteen regions, and the four exceptions were Uiseong, Haman, Buyeo, and Iksan regions. Also, BWYV-infected weeds [i.e., Oriental smartweed (Persicaria blumei), Thunberg’s smartweed (Persicaria thunbergii), Asian copperleaf (Acalypha australis), Pigeo (Phytolacca americana), Indian lettuce (Lactuca indica), Edible amaranth (Amaranthus inamoenus), and Ruderal chaff flower (Achyranthes fauriei H.Lév. & Vaniot) were reported in Jeonnam Province by Dr. Sug-Ju Ko (not published). From 2015 to 2016, the occurrence pattern of the viruses on pepper in fields showed that BWYV had the third highest incidence among the seven viruses (Kwon et al., 2018). These research reports showed that BWYV exists in Solanaceae crops (hot pepper, bell pepper, and paprika) and various weeds on a national scale in Korea. These results imply that BWYV has already spread to various weeds or the virus has become endemic in agricultural ecosystems in Korea.
Sequence properties of BWYV and phylogenetic analysis
Four specific primer sets were designed to determine the complete genomic sequence of the BWYV-Korea isolate. The sequence of the BWYV-Korea genome was assembled from four overlapping fragments. The complete genomic sequence of the BWYV-Korea isolate comprised 5750 nucleotides (nt), including 28 nt at the 5’-terminal UTR and 203 nt at the 3’-terminal UTR. An intergenic UTR of 200 nt (nt 3332-3531) was confirmed between ORF 2 and ORF 3. The BWYV-Korea isolate has six ORFs whose start codon and termination codon nucleotide positions are as follows: ORF0 (P0 protein: 29-31/743-745), ORF1 (P1 protein: 150-152/2055-2057), ORF2 (P1-P2 protein: 150-152/3329-3331), ORF3 (P3 protein: 3532-3534/4138-4140), ORF4 (P4 protein: 3563-3565/4088-4090), and ORF5 (P3-P5 protein: 3532-3534/5545-5547). The ORF2 overlaps with ORF1 and encodes the P1-P2 fusion protein by a -1 ribosomal frameshift at nucleotide positions 1454 to 1456. The ORF5 encodes the P3-P5 fusion protein with a read-through of the P3 stop codon at nucleotide positions 4140 to 4142. The conserved protein domains of the deduced proteins of the BWYV-Korea isolate were analyzed using BLASTX searches. It contained Luteovirus P0 protein domain, which is a viral suppressor of gene silencing in P0, a serine-like protease (peptidase S39) in P1, a RNA-dependent RNA polymerase in P2, Luteovirus coat protein domain in P3, movement protein in P4, and translational read-through protein domain in P3-P5. The first eight nucleotides of the conserved motif (ACAAAAGA) was confirmed at the 5’ terminus, which exists in all the polerovirus species (Knierim et al., 2013). The genome organization of the BWYV-Korea isolate is similar to another BWYV isolate as shown in Fig. 4. The complete genome sequence of the BWYV-Korea isolate has been deposited in NCBI GenBank under the accession number LC198684.

The predicted genomic organization of the BWYV-Korea isolate. The complete genome sequence of the BWYV-Korea isolate comprises 5750 nucleotides (nt). ORF2 overlaps with ORF1 and encodes the P1–P2 fusion protein with a -1 ribosomal frameshift at nt 1454 (left arrow). ORF5 encodes the P3–P5 protein with a read-through of the P3 stop codon at position 4140 (right arrow).
The complete genome of the BWYV-Korea isolate shares a nucleotide identity ranging from 80.6-97.1% with the other 7 BWYV isolates. The deduced amino acid sequences of BWYV-Korea isolate had amino acid sequence identities ranging from 82.4-90.3% for P0, 84.8-96.7% for P1, 90.8-97.5% for P2, 87.1-100% for P3, 78.9-96.6% for P4, and 60.2-99.3% for P5 with other reported BWYV isolates (Table 3). The complete nucleotide sequence and all the ORFs (P0, P1, P1-P2, P3, P4 and P3-P5) showed the highest nucleotide and amino acid identity with the BWYV LS isolate. This LS isolate was identified from a weed (Leonurus sibiricus) in South Korea, and its 3’-terminal UTR was 10-nt shorter than the BWYV Korea isolate (Kwon et al., 2016).
Percentage identity of the complete nucleotide (nt) sequence and deduced amino acid (aa) sequence between BWYV-Korea isolate and previously reported BWYV isolates
The phylogenetic tree was constructed to determine the relatedness between the BWYV-Korea isolate and other BWYV isolates, including two species as outgroup. An analysis was conducted with the complete nucleotide sequence and P3 (coat protein) amino acid sequence (Fig. 5). The phylogenetic tree showed that the BWYV-Korea isolate was closely related to the BWYV LS isolate.

Phylogenetic trees generated by the alignment of the complete nucleotide genome and deduced amino acid sequence with other BWYV isolates and outgroup (BYDV-PAV and PEMV-1). (A) complete genome, (B) P3 protein. The abbreviations of virus name and GenBank accession number is as follows: BWYV isolates (BJ: HM804472, Korea: LC198684, LS: KM076647, Mulhouse: KU521324, Rouen1: KU521325, Rouen2: KU521326, USA: AF473561), BYDV-PAV (Barley yellow dwarf virus–PAV, EF521849), and PEMV-1 (Pea enation mosaic virus 1, NC003629). The bootstrap analysis was carried out using 1000 replicates and scale bar represents 0.05 nucleotide substitutions per site.
BWYV is an emerging virus in Korea, spread by insect vectors. It is thought that some weeds also play the role of a reservoir host. To reduce the damage caused by BWYV, it is necessary to remove the insect vector and susceptible weeds. To our best knowledge, this is the first report of BWYV infection of paprika plants and the complete genome sequencing of the infectious agent.
Acknowledgement
This work was carried out with the support of “Cooperative Research Program for Agriculture Science and Technology Development (Project No. PJ013741)” Rural Development Administration, Republic of Korea.
References
Notes
Conflicts of Interest
No potential conflict of interest relevant to this article was reported.