ABSTRACT
In April 2014, seedlings of Fallopia multiflora showing wilt symptom were first found at a greenhouse in Punggi-eup, Yeongju-si, Korea. A Fusarium-like fungus was isolated from the wilted plant and it was identified as Fusarium oxysporum based on morphological characteristics and nucleotide sequence data of translation elongation factor 1-α. The fungus isolated from the diseased plant was revealed to be pathogenic to the host plant through pathogenicity tests, and the reisolation of the pathogen confirmed Koch’s postulates. This is the first report of Fusarium wilt occurring on Fallopia multiflora in the world.
Fallopia multiflora (Thunb.) Haraldson (syn.
Polygonum multiflorum Thunb., family Polygonaceae), commonly called hasuo in Korean, has been commercially cultivated as a medicinal plant in Korea. In particular, tuberous roots of
F. multiflora (Polygoni Multiflori Radix) have been used for traditional medicine in China, Japan, and Korea. In April 2014, symptoms of wilting were observed on
F. multiflora seedlings, with disease incidence of up to 30%, at a nursery located in Punggi-eup, Yeongju-si, Korea (
Fig. 1A and
1B). The initial symptoms appeared as brown discoloration on the lower part of stems (
Fig. 1C,
Fig. 1 left). The affected young seedlings were wilted with sudden drooping, followed by drying of leaves. At advanced stages of the disease, severely infected plants were collapsed and later covered by whitish fungal mass (
Fig. 1C).
Fig. 1
Wilt symptoms caused by Fusarium oxysporum on Fallopia multiflora seedlings. A, B: Plants affected by wilt disease in a seedling nursery (red arrow indicates the wilted plant), C: A diseased plant showing brown discoloration of the lower stem (left) and a decayed plant overgrown by the fungal pathogen (right).

Small pieces of the diseased plant tissues were sterilized with 70% ethanol for 1 min followed by 2 washes of sterile water. They were placed onto Petri dishes containing potato dextrose agar (PDA, Difco, USA), and incubated at room temperature. After 2 days, without bacterial or fungal contaminants, hyphal tips of the emerging fungus were transferred to new PDA plate to obtain pure culture. A
Fusarium-like isolate 14-147 was obtained and identified based on morphology and phylogenetic analysis of partial translation elongation factor-1a (
TEF). Morphological features of the fungus grown on PDA for 10 days were observed under a Zeiss AXIO Imager A1 microscope. The fungus produced whitish cottony colony with the aerial mycelia becoming tinged in purple on PDA. The colony on PDA reached the edge of the plate after 10 days of incubation at 25°C under continuous fluorescent light (
Fig. 2A). Macroconidia were mostly three septate, falcate, with foot-shaped basal cell, and 30-63 × 8.5-13.5 mm in size (
Fig. 2B). Abundant microconidia were formed on monophialides, and 7-16 × 3-5 mm in size (
Fig. 2C). Chlamydospores were terminal or intercalary in position and 6-13 mm in size (
Fig. 2D). On the basis of the morphological characteristics, the fungus matched well with the description of
Fusarium oxysporum (
Samson et al., 2004).
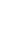
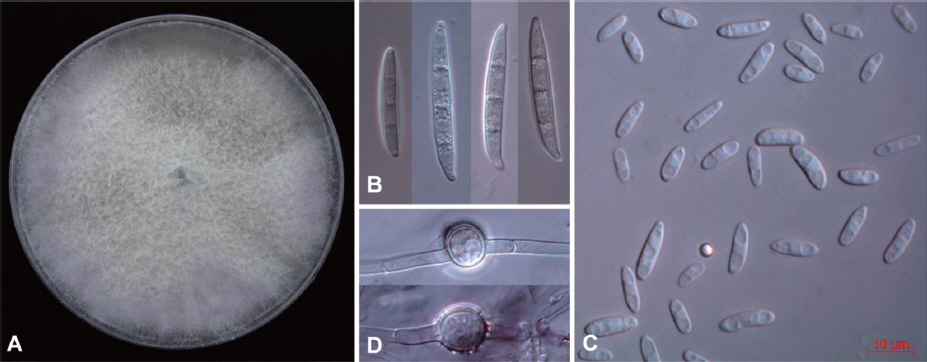
Fig. 2
Cultural and morphological characteristics of Fusarium oxysporum 14-147 from Fallopia multiflora seedlings. A: Colony on potato dextrose agar after 10 days of incubation at room temperature, B: Macroconidia, C: Micoroconidia, D: Chlamydospores. Scale bar: 10 mm (B, C and D).
Genomic DNA was extracted from fungal mycelia using DNeasy plant mini kit (Qiagen, USA). The
TEF gene region was amplified using the primer pairs EF1 (5’-ATG GGT AAG GAR GAC AAG AC-3’) and EF2 (5’-GGA RGT ACC AGT SAT CAT GTT-3’) (
O’Donnell et al., 1998). The PCR poducts were purified and directly sequenced with the same primers. The raw sequence of
F. oxysporum was assembled and edited using SeqMan (DNASTAR Lasergene, Madison, WI, USA). The sequence data obtained was compared with all fungal sequences available in the GenBank database using BLAST search tool. The BLAST search showed that the present sequence was 99% similar to the
TEF sequences of
F. oxysporum. To infer the phylogenetic relationship, 11 reference sequences of
Fusarium spp. were taken from GenBank. Two sequences of
Fusarium solani (JF740866/JX118990) were used to root the tree. A neighbor-joining tree (
Fig. 3) was constructed with the Tajima-Nei model using MEGA5 (
Tamura et al., 2011). The sequence of the present isolate clustered with those of
F. oxysporum, forming a distinct group with a bootstrap value of 100%. Accordingly, the molecular analysis confirmed the morphological identification for the fungal pathogen.
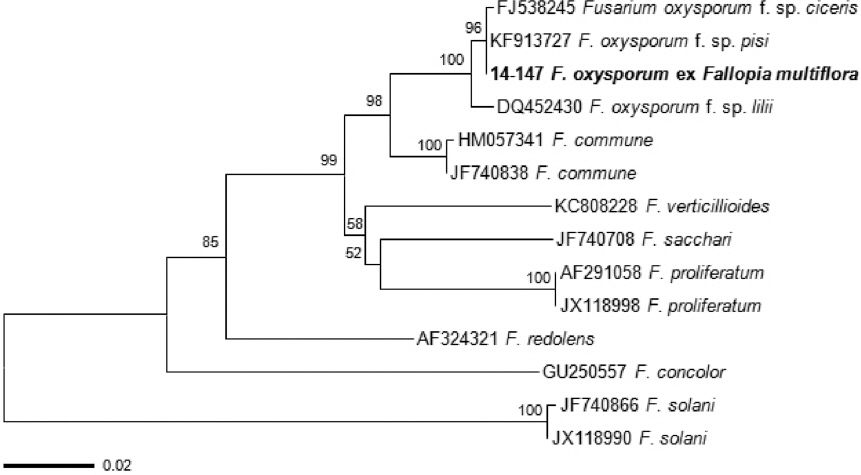
Fig. 3
Phylogenetic tree showing the relationship between Fusarium oxysporum isolate 14-147 and other Fusarium isolates based on translation elongation factor-1α.
Koch’s postulates were demonstrated for the present isolate by dipping the roots of 2-month-old seedlings in a spore suspension (1 × 106 conidia/ml) for 10 min. The inoculated plants were placed in a growth chamber maintained at 25°C under an alternate 12 hr photoperiod. Control plants were sprayed with sterile distilled water using the same method. The plants were covered with plastic bags and kept under laboratory conditions at 25°C. After 48 hr, they were then placed under natural conditions. Any disease symptoms were not shown on the control plants two weeks after inoculation. In contrast, all inoculated plants exhibited vascular discoloration, wilting of leaves, followed by collapse of the plant. The fungus was successfully re-isolated from the symptomatic plants.
F. oxysporum is well-known as one of the most important soil-borne fungal pathogens. There have been no reports of the occurrence of the wilt disease caused by
F. oxysporum on
F. multiflora. According to the Systematic Mycology and Microbiology Laboratory Fungus-Host Database, one species of
Fusarium,
F. oidioides, was reported on
F. multiflora (
Farr and Rossman, 2014). To our knowledge, this is the first report of
F. oxysporum associated with
F. multiflora seedlings in Korea. The finding of the disease on
F. multiflora in a seedling greenhouse indicates that Fusarium wilt caused by
F. oxysporum can pose a potential threat to nursery seedling production for the plants in Korea.
Acknowledgements
We would like to thank Dr. Seung-Beom Hong (KACC, NAAS, RDA) for providing valuable opportunities for DIC microscopy. This work was carried out with the support of ‘‘Cooperative Research Program for Agriculture Science & Technology Development (Project No. PJ01136802)’’ Rural Development Administration, Republic of Korea.
References
O'Donnell, K, Kistler, H. C, Cigelnik, E and Ploetz, R. C Multiple evolutionary origins of the fungus causing Panama disease of banana: concordant evidence from nuclear and mitochondrial gene genealogies.
Proc. Natl. Acad. Sci. USA 1998. 95: 2044-2049.

Samson, R. A, Hoekstra, E. S and Frisvad, J. C Introduction to Food- and Airborne Fungi. 2004. 7th ed The Netherlands. Centraalbureau voor Schimmelcultures, pp. 389 pp.
Tamura, K, Peterson, D, Peterson, N, Stecher, G, Nei, M and Kumar, S MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods.
Mol. Biol. Evol 2011. 28: 2731-2739.












